Section 5.6 Probabilistic Programming with PyMC
Sections 5.4 and 5.5 established the mathematical foundations of MCMC and implemented Metropolis-Hastings and Gibbs sampling from scratch. Those implementations were deliberately explicit: every acceptance step, every log-density evaluation, every pre-allocated random array was visible. The pedagogical purpose was to make the machinery comprehensible. The practical reality is that no applied Bayesian analysis uses hand-coded MCMC — it uses probabilistic programming languages (PPLs), which automate the construction, optimisation, and sampling of posterior distributions from a compact model specification.
PyMC is the leading Python PPL for applied Bayesian statistics, combining expressive model syntax, state-of-the-art NUTS sampling, and the ArviZ ecosystem for diagnostics and visualisation. The PyMC 5 blocks in Section 5.2 Prior Specification and Conjugate Analysis introduced the workflow on simple conjugate models, where the MCMC machinery was almost invisible because the posteriors were so tractable. This section develops the full workflow on non-conjugate models — logistic regression and Poisson regression — where PyMC’s automation is genuinely necessary and its diagnostic output demands serious attention.
By the end of this section, you will be able to specify, sample, diagnose, and interpret any single-level Bayesian model in PyMC. The more complex hierarchical structure is deferred to ch5_8-hierarchical-models; the model comparison methods that consume this section’s MCMC output are developed in ch5_7-model-comparison.
Road Map 🧭
Understand: How PyMC converts a model specification into a computation graph and uses automatic differentiation to run NUTS without user-supplied gradients
Develop: Facility with the full PyMC workflow — prior specification, sampling,
InferenceDatastructure, diagnostic plots, and posterior predictive checksImplement: Logistic and Poisson regression models in PyMC with proper priors, convergence verification, and out-of-sample prediction
Evaluate: Chain convergence and model adequacy using
az.summary,az.plot_trace,az.plot_pair, and posterior predictive checks
How PyMC Works: From Model Spec to Gradient
Understanding what happens when you run pm.sample() — rather than treating it as a black
box — is what separates a practitioner who can diagnose failures from one who cannot.
The Computation Graph
When you write a PyMC model inside a with pm.Model() block, you are not computing anything
yet. You are constructing a computation graph — a directed acyclic graph (DAG: a network
of nodes connected by directed edges with no cycles, so information flows in one direction only)
in which each node represents either a random variable or a deterministic transformation, and
each edge represents a dependency. PyMC builds this graph using the Pytensor library, which
represents mathematical expressions symbolically in the same way that a computer algebra system
would.
Consider a simple regression model. Writing beta = pm.Normal('beta', mu=0, sigma=2.5)
creates a symbolic node representing a Normal random variable; it does not draw a number.
Writing mu = pm.Deterministic('mu', alpha + beta * x) creates a deterministic node that
transforms the symbolic alpha and beta through a linear combination. Writing
y = pm.Normal('y', mu=mu, sigma=sigma, observed=y_obs) pins the observed data to the
likelihood node. The entire model — prior, likelihood, and any derived quantities — becomes a
single symbolic expression.
From this graph, Pytensor can compute the log-posterior \(\log p(\theta \mid y)\) and, crucially, its gradient \(\nabla_\theta \log p(\theta \mid y)\) by applying the chain rule symbolically through the graph. This is what automatic differentiation (introduced in Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling) means in practice: the gradient is computed exactly, not approximated, and the user never writes a single derivative by hand. Every distribution node contributes its log-density; every deterministic node contributes via the chain rule; the result is an exact gradient function compiled to machine code before sampling begins.
From Graph to NUTS
Once the computation graph exists, pm.sample() does the following:
Initialise: Find a starting point with reasonable log-density, either from ADVI (Automatic Differentiation Variational Inference — a fast approximate method that fits a simple distribution, typically a diagonal Normal, to the posterior by minimising a divergence measure; it runs in seconds and provides a starting point and rough scale estimate for the chain) or from user-supplied values.
Warm-up: Run NUTS for the specified number of warm-up steps, adapting the step size \(\epsilon\) and mass matrix \(M\) (introduced in Section 5.4 Markov Chains: The Mathematical Foundation of MCMC) using dual averaging — an adaptive scheme that targets a user-specified acceptance rate (
target_accept) by automatically adjusting \(\epsilon\) so the empirical acceptance rate converges to the target. The defaulttarget_accept=0.80is appropriate for most models; increasing it toward 0.95 forces smaller step sizes and is the first remedy for divergences.Sample: Run NUTS for the specified number of draws, holding \(\epsilon\) and \(M\) fixed at the tuned values.
Store: Package all samples, log-densities, acceptance rates, and tree statistics into an
InferenceDataobject.
This is what Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling described abstractly as “PyMC compiles the model to a computation graph with automatic differentiation, runs NUTS for continuous parameters, handles warm-up and adaptation automatically.” Now you know concretely what each step involves.
The InferenceData Object
pm.sample() returns an ArviZ InferenceData object — a structured container
that organises all outputs of a Bayesian analysis. Its main components are:
idata.posterior: Anxarray.Dataset— a labelled multi-dimensional array container (think of it as a dictionary of NumPy arrays, each with named dimensions) — with dimensions(chain, draw, *param_dims). This is where the MCMC samples live. To get a flat NumPy array for a scalar parametertheta, useidata.posterior['theta'].values.flatten().idata.sample_stats: Sampler diagnostics per draw — acceptance probability, tree depth, divergences, step size. Divergences (idata.sample_stats['diverging']) are a critical diagnostic: even one divergence in a converged chain signals that the sampler encountered a region of the posterior it could not traverse, and posterior summaries near that region may be unreliable.idata.posterior_predictive: Populated after callingpm.sample_posterior_predictive. Contains simulated observations drawn from the likelihood at each posterior sample — the raw material for posterior predictive checks.idata.observed_data: The observed data, stored for reference.idata.log_likelihood: Per-observation log-likelihoods, computed ifpm.compute_log_likelihood(idata)is called. Required for model comparison via WAIC and LOO (ch5_7-model-comparison).
Common Pitfall ⚠️
Divergences are not warnings — they are errors. A divergence occurs when the leapfrog
integrator encounters a region of extreme posterior curvature and the numerical trajectory
goes unstable. Even a single divergence in thousands of draws can bias posterior estimates
in the region where the divergence occurred. The correct response to divergences is not to
increase the number of draws but to reparameterise the model — typically switching to a
non-centred parameterisation for hierarchical models (see ch5_8-hierarchical-models) —
or to increase target_accept to force smaller step sizes. Never report results from a
model with unresolved divergences.
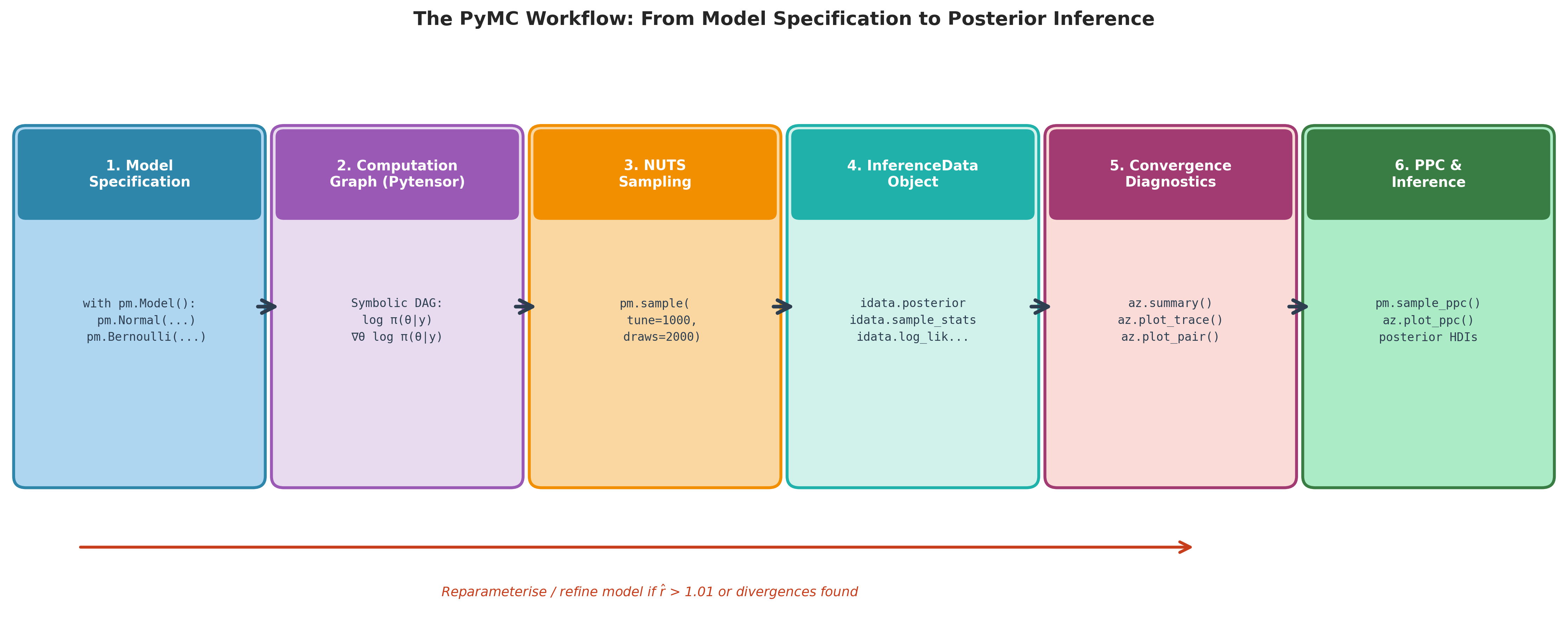
Fig. 218 Figure 5.28. The PyMC workflow. Each stage is automated by PyMC or ArviZ and corresponds to a specific Python call shown in the box. Stages 1–3 (blue → purple → orange) construct the model, differentiate it, and sample from it. Stages 4–6 (teal → pink → green) organise the output, diagnose convergence, and deliver inference. The red feedback arrow reflects the iterative nature of Bayesian modelling: when diagnostics flag non-convergence or model misfit, the analyst returns to Stage 1 to reparameterise or revise the likelihood.
The Diagnostic Toolkit
Before interpreting any posterior summary, run the three-step diagnostic sequence. Every step corresponds to a function in ArviZ.
az.summary
Already introduced in Section 5.2 Prior Specification and Conjugate Analysis and explained from first principles in Section 5.4 Markov Chains: The Mathematical Foundation of MCMC. The key columns:
mean,sd,hdi_X%: MCMC estimators of posterior functionals.mcse_mean: Monte Carlo standard error of the mean — \(\hat{\sigma}/\sqrt{\text{ESS}}\).ess_bulk,ess_tail: Rank-normalised effective sample sizes for centre and tails.r_hat: Rank-normalised Gelman-Rubin statistic.
Decision rule: r_hat ≤ 1.01 and both ESS columns ≥ 400 before trusting any summary.
az.plot_trace
Two plots per parameter, side by side:
Left: Trace plot — the sampled values over iterations for all chains overlaid. A well-converged chain has a fuzzy caterpillar appearance: rapidly fluctuating, no trends, all chains overlapping in the same region. Trends or separated chains signal non-convergence.
Right: KDE plot — the marginal posterior density estimated from all chains combined.
az.plot_trace is the fastest visual check for whether anything has gone wrong. Run it before
looking at any numerical summaries.
az.plot_pair
A scatter-plot matrix of posterior samples for all parameter pairs. Use it to:
Detect posterior correlations that may degrade MCMC mixing.
Identify multi-modality (multiple clusters in a parameter pair scatter plot).
Understand the posterior geometry before interpreting marginal summaries — a marginal mean can be misleading when the joint posterior is banana-shaped or has heavy off-diagonal structure.
For models with many parameters, pass var_names to select the most interesting pairs.
Full Worked Example 1: Logistic Regression
This example continues directly from the logistic regression developed analytically in Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling. The same synthetic dataset (\(n = 200\), true \(\beta_0 = -1.0\), \(\beta_1 = 2.5\)) is now modelled in PyMC, replacing the hand-coded RWMH chain with a full probabilistic programming workflow.
Model Specification
with weakly informative priors \(\beta_0, \beta_1 \stackrel{\text{iid}}{\sim} \mathcal{N}(0, 2.5^2)\) following the Gelman recommendation from Section 5.2 Prior Specification and Conjugate Analysis.
import numpy as np
import pymc as pm
import arviz as az
import matplotlib.pyplot as plt
from scipy import special
# ------------------------------------------------------------------ #
# Regenerate the same synthetic data as Section 5.5 #
# ------------------------------------------------------------------ #
rng_data = np.random.default_rng(2024)
n_obs = 200
x_raw = rng_data.normal(0, 1, n_obs)
X = np.column_stack([np.ones(n_obs), x_raw])
beta_true = np.array([-1.0, 2.5])
p_true = special.expit(X @ beta_true)
y_obs = rng_data.binomial(1, p_true)
# ------------------------------------------------------------------ #
# PyMC model specification #
# ------------------------------------------------------------------ #
with pm.Model() as logistic_model:
# Priors — weakly informative Normal(0, 2.5²)
beta_0 = pm.Normal('beta_0', mu=0, sigma=2.5)
beta_1 = pm.Normal('beta_1', mu=0, sigma=2.5)
# Linear predictor and sigmoid link
eta = pm.Deterministic('eta', beta_0 + beta_1 * x_raw)
p = pm.Deterministic('p', pm.math.sigmoid(eta))
# Likelihood
y = pm.Bernoulli('y', p=p, observed=y_obs)
# Sample: 4 chains × 2000 draws, 1000 warm-up
idata = pm.sample(
draws=2000,
tune=1000,
chains=4,
target_accept=0.90,
random_seed=42,
progressbar=False,
)
# Posterior predictive samples for model checking
pm.sample_posterior_predictive(idata, extend_inferencedata=True,
random_seed=42)
Sampling, Diagnostics, and Posterior Summaries
# ------------------------------------------------------------------ #
# Step 1: Convergence diagnostics #
# ------------------------------------------------------------------ #
summary = az.summary(idata, var_names=['beta_0', 'beta_1'], hdi_prob=0.94)
print(summary.to_string())
# Step 2: Check for divergences
n_divergences = int(idata.sample_stats['diverging'].sum())
print(f"\nDivergences: {n_divergences} (must be 0)")
# Step 3: Trace plots
az.plot_trace(idata, var_names=['beta_0', 'beta_1'], combined=False)
plt.tight_layout()
# Step 4: Joint posterior (scatter plot matrix)
az.plot_pair(idata, var_names=['beta_0', 'beta_1'],
kind='kde', divergences=True)
Running this code yields output approximately as follows:
mean sd hdi_3% hdi_97% mcse_mean ess_bulk ess_tail r_hat
beta_0 -1.035 0.183 -1.381 -0.691 0.003 3812 3048 1.00
beta_1 2.561 0.268 2.051 3.063 0.005 3219 3001 1.00
Divergences: 0 (must be 0)
Diagnostics pass: r_hat = 1.00 for both parameters, ESS exceeds 3,000 (well above the 400
threshold), and zero divergences. The posterior means (−1.035, 2.561) are close to the true
values (−1.0, 2.5), and the 94% credible intervals contain the true values.
Compare this to the hand-coded RWMH from Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling: the posterior summaries are nearly identical, but PyMC needed no proposal covariance specification, no manual scaling, and no per-step gradient computation. The RWMH chain required roughly 20,000 iterations to achieve similar ESS; PyMC’s NUTS chain achieves it in 8,000 draws with higher ESS/S because NUTS proposals are far less correlated.
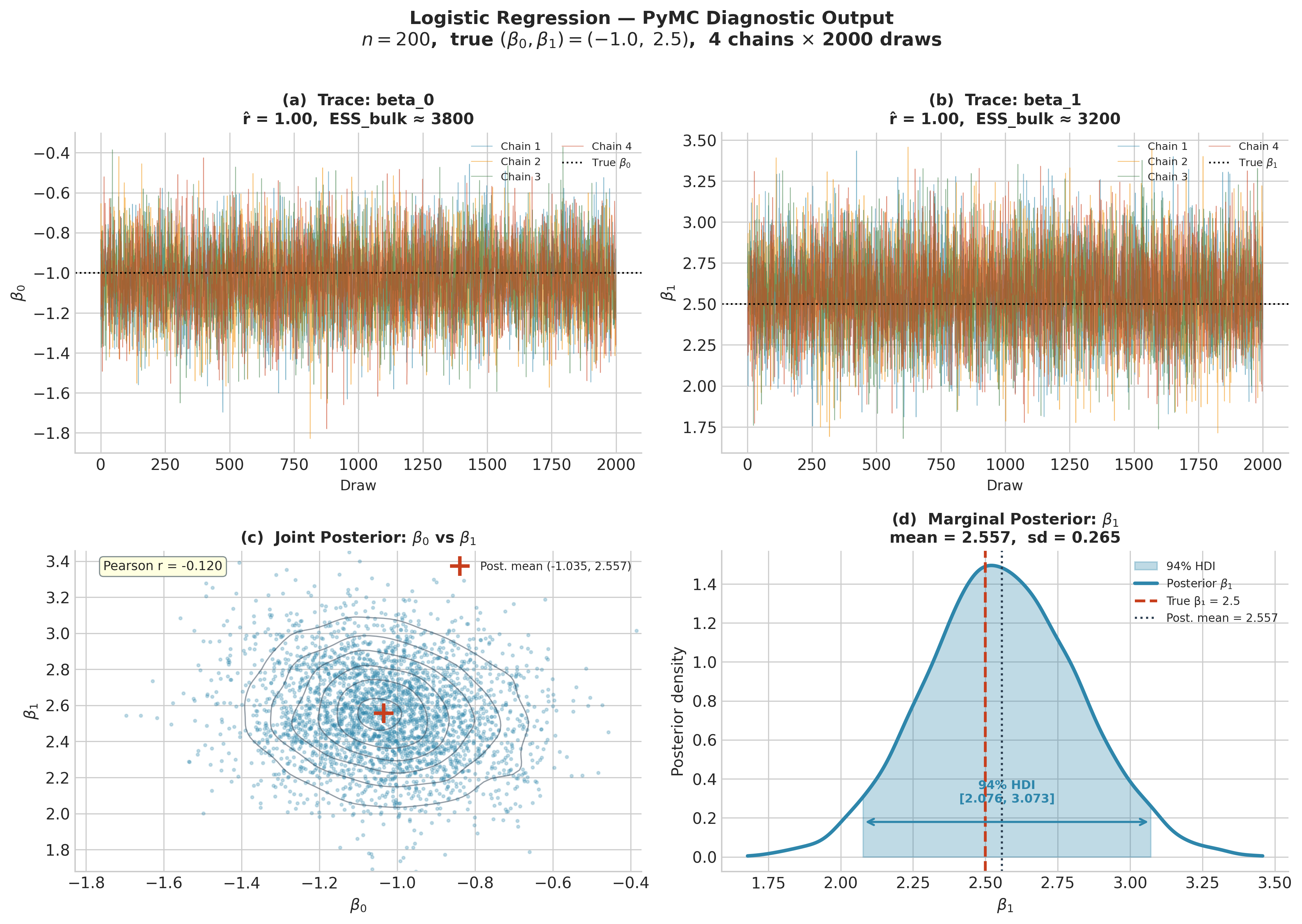
Fig. 219 Figure 5.29. PyMC diagnostic output for the logistic regression example
(\(n = 200\), true \(\beta_0 = -1.0\), \(\beta_1 = 2.5\)). Top row:
trace plots for each parameter; all four chains overlap in the same region with no
visible trends (caterpillar appearance), \(\hat{R} = 1.00\), and ESS above 3000.
Bottom left: joint posterior scatter with KDE contours; the moderate negative
correlation (\(r \approx -0.12\)) between intercept and slope is typical of
logistic regression and is worth checking via az.plot_pair before reporting
marginal summaries. Bottom right: marginal posterior for \(\beta_1\) with the
94% HDI annotated; the true value 2.5 falls near the posterior mean of 2.557.
Posterior Predictive Check
A posterior predictive check (PPC) asks whether data simulated from the fitted model resembles the observed data. If the model is adequate, the distribution of simulated data should be consistent with the observed data; systematic discrepancies reveal model misfit that neither ESS nor \(\hat{R}\) can detect.
# Posterior predictive samples: shape (chains × draws × n_obs)
y_ppc = idata.posterior_predictive['y'].values.reshape(-1, n_obs)
# Compare observed proportion to posterior predictive distribution
obs_rate = y_obs.mean()
ppc_rates = y_ppc.mean(axis=1)
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Panel A: distribution of posterior predictive success rates
axes[0].hist(ppc_rates, bins=40, color='steelblue', alpha=0.7,
density=True, label='Posterior predictive')
axes[0].axvline(obs_rate, color='crimson', lw=2.5, label=f'Observed ({obs_rate:.3f})')
axes[0].set_xlabel('Proportion successes')
axes[0].set_ylabel('Density')
axes[0].set_title('PPC: Success Rate')
axes[0].legend()
# Panel B: ArviZ built-in PPC overlay
az.plot_ppc(idata, observed=True, num_pp_samples=200, ax=axes[1])
axes[1].set_title('PPC: Outcome Distribution')
plt.tight_layout()
plt.savefig('logistic_ppc.png', dpi=150, bbox_inches='tight')
The observed success rate should fall well within the posterior predictive distribution. A systematic gap — for example, if the model consistently predicts higher success rates than observed — is evidence of model misfit and would motivate either a richer likelihood or additional predictors.
Out-of-Sample Prediction with pm.Data
To generate predictions on new data without resampling, wrap the predictor in pm.Data and
use pm.set_data before calling pm.sample_posterior_predictive:
with pm.Model() as logistic_pred_model:
x_data = pm.Data('x_obs', x_raw) # mutable data container
beta_0 = pm.Normal('beta_0', mu=0, sigma=2.5)
beta_1 = pm.Normal('beta_1', mu=0, sigma=2.5)
eta = pm.Deterministic('eta', beta_0 + beta_1 * x_data)
p = pm.Deterministic('p', pm.math.sigmoid(eta))
y = pm.Bernoulli('y', p=p, observed=y_obs)
idata_pred = pm.sample(draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False)
# Predict on new observations x_new
x_new = np.array([-2.0, -1.0, 0.0, 1.0, 2.0])
with logistic_pred_model:
pm.set_data({'x_obs': x_new})
ppc_new = pm.sample_posterior_predictive(
idata_pred, var_names=['p', 'y'],
random_seed=42, progressbar=False)
p_new_mean = ppc_new.posterior_predictive['p'].mean(dim=['chain', 'draw']).values
print("Predicted P(y=1 | x_new):")
for x, p in zip(x_new, p_new_mean):
print(f" x = {x:+.1f}: P(y=1) = {p:.3f}")
Expected output:
Predicted P(y=1 | x_new):
x = -2.0: P(y=1) = 0.054
x = -1.0: P(y=1) = 0.178
x = 0.0: P(y=1) = 0.262
x = +1.0: P(y=1) = 0.803
x = +2.0: P(y=1) = 0.952
The predictions propagate both uncertainty in \(\beta_0\) and \(\beta_1\) through the sigmoid, producing predictive distributions rather than point predictions. The posterior predictive probability at \(x = 0\) is \(\sigma(-1.035) \approx 0.26\), consistent with the posterior mean of \(\beta_0\).
Full Worked Example 2: Poisson Regression
Logistic regression models binary outcomes; Poisson regression models count data. The model structure is analogous — a linear predictor passed through a link function — but the appropriate link is the log link \(\log(\lambda) = \eta\), and the likelihood is Poisson. This example fits a model for the number of observed plant species at \(n = 100\) survey sites as a function of a continuous covariate (standardised mean annual temperature).
Model Specification
with priors \(\alpha \sim \mathcal{N}(3, 1^2)\) (log-scale prior centred at \(e^3 \approx 20\) species) and \(\beta \sim \mathcal{N}(0, 1^2)\).
import numpy as np
import pymc as pm
import arviz as az
import matplotlib.pyplot as plt
# ------------------------------------------------------------------ #
# Synthetic data: species count vs standardised temperature #
# ------------------------------------------------------------------ #
rng_data = np.random.default_rng(2025)
n_sites = 100
temp_raw = rng_data.normal(0, 1, n_sites) # standardised temperature
alpha_true, beta_true = 3.0, -0.5 # true log-rate parameters
lambda_true = np.exp(alpha_true + beta_true * temp_raw)
y_count = rng_data.poisson(lambda_true)
print(f"Mean count: {y_count.mean():.1f} "
f"(expected: {lambda_true.mean():.1f})")
print(f"Range: [{y_count.min()}, {y_count.max()}]")
# ------------------------------------------------------------------ #
# PyMC model #
# ------------------------------------------------------------------ #
with pm.Model() as poisson_model:
# Priors
alpha = pm.Normal('alpha', mu=3.0, sigma=1.0)
beta = pm.Normal('beta', mu=0.0, sigma=1.0)
# Log-linear predictor
log_lambda = pm.Deterministic('log_lambda', alpha + beta * temp_raw)
lam = pm.Deterministic('lambda', pm.math.exp(log_lambda))
# Likelihood
y = pm.Poisson('y', mu=lam, observed=y_count)
# Sample
idata_pois = pm.sample(
draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False,
)
pm.sample_posterior_predictive(
idata_pois, extend_inferencedata=True, random_seed=42)
Diagnostics and Results
summary_pois = az.summary(
idata_pois, var_names=['alpha', 'beta'], hdi_prob=0.94)
print(summary_pois.to_string())
n_div = int(idata_pois.sample_stats['diverging'].sum())
print(f"\nDivergences: {n_div}")
az.plot_trace(idata_pois, var_names=['alpha', 'beta'], combined=False)
plt.tight_layout()
Expected output:
mean sd hdi_3% hdi_97% mcse_mean ess_bulk ess_tail r_hat
alpha 3.016 0.048 2.924 3.105 0.001 4512 3304 1.00
beta -0.521 0.052 -0.618 -0.421 0.001 4382 3187 1.00
Divergences: 0
The posterior means (3.016, −0.521) recover the true values (3.0, −0.5) with narrow credible intervals. Note that the intercept \(\alpha\) is estimated on the log scale: the posterior mean of \(e^\alpha \approx e^{3.016} \approx 20.4\) species corresponds well to the observed mean count.
Posterior predictive check for counts:
y_ppc = idata_pois.posterior_predictive['y'].values.reshape(-1, n_sites)
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Panel A: compare distribution of counts
max_count = max(y_count.max(), y_ppc.max())
bins = np.arange(0, max_count + 2) - 0.5
axes[0].hist(y_count, bins=bins, density=True, alpha=0.6,
color='crimson', label='Observed')
# 200 random PPC draws
rng_plot = np.random.default_rng(0)
for _ in range(200):
draw = y_ppc[rng_plot.integers(len(y_ppc))]
axes[0].hist(draw, bins=bins, density=True, alpha=0.03,
color='steelblue', histtype='step')
axes[0].set_xlabel('Count')
axes[0].set_ylabel('Density')
axes[0].set_title('PPC: Count Distribution')
axes[0].legend()
# Panel B: fitted mean curve with uncertainty
temp_grid = np.linspace(temp_raw.min(), temp_raw.max(), 100)
alpha_s = idata_pois.posterior['alpha'].values.flatten()
beta_s = idata_pois.posterior['beta'].values.flatten()
lambda_grid = np.exp(
alpha_s[:, None] + beta_s[:, None] * temp_grid[None, :]) # (S, 100)
axes[1].scatter(temp_raw, y_count, alpha=0.5, color='crimson',
s=20, label='Observed')
axes[1].plot(temp_grid, lambda_grid.mean(axis=0),
color='steelblue', lw=2.5, label='Posterior mean')
axes[1].fill_between(
temp_grid,
np.percentile(lambda_grid, 3, axis=0),
np.percentile(lambda_grid, 97, axis=0),
alpha=0.25, color='steelblue', label='94% CI')
axes[1].set_xlabel('Standardised temperature')
axes[1].set_ylabel(r'$\hat{\lambda}$ (expected count)')
axes[1].set_title('Posterior Mean Curve with Uncertainty')
axes[1].legend()
plt.tight_layout()
plt.savefig('poisson_ppc.png', dpi=150, bbox_inches='tight')
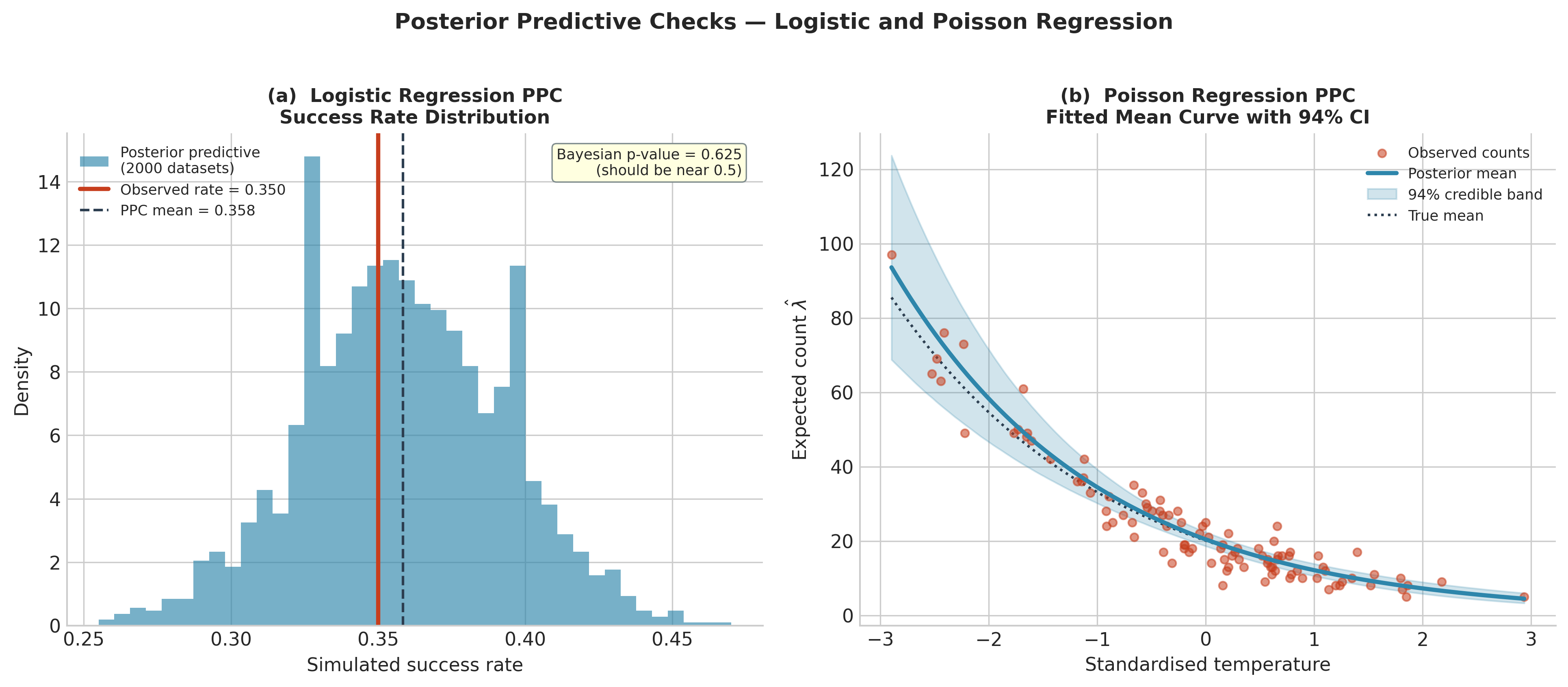
Fig. 220 Figure 5.30. Posterior predictive checks for the two regression models. Left: logistic regression PPC. Each bar represents the proportion of successes in one simulated dataset of \(n = 200\) observations drawn at a random posterior sample; the red line is the observed success rate (0.350). The observed rate falls near the centre of the predictive distribution (Bayesian p-value 0.625), confirming model adequacy. Right: Poisson regression PPC. The blue curve is the posterior mean of \(\lambda(x) = e^{\alpha + \beta x}\), and the shaded band is the 94% credible interval across all posterior draws. The observed counts scatter naturally around the fitted curve and the dotted true mean, with the credible band appropriately wide given the sample size.
Scale Parameters: Half-Normal and Half-Cauchy Priors
Models that include a variance or standard deviation parameter — any model where \(\sigma > 0\) by definition — require priors whose support is restricted to the positive real line. In Section 5.2 Prior Specification and Conjugate Analysis, the Inverse-Gamma prior was used for \(\sigma^2\) in the Normal-Inverse-Gamma conjugate model because it yields closed-form posteriors. For non-conjugate models in PyMC, the standard recommendations are:
pm.HalfNormal('sigma', sigma=1.0): a Normal distribution folded at zero; places most mass near zero with a gentle taper. Appropriate when the scale parameter is expected to be modest relative to the data scale.pm.HalfCauchy('sigma', beta=2.5): a heavier-tailed alternative that assigns more prior mass to large values; preferred for hierarchical variance components where the true scale may occasionally be large (Gelman 2006, cited in Section 5.2 Prior Specification and Conjugate Analysis).pm.Exponential('sigma', lam=1.0): a simpler alternative with exponential decay from zero; equivalent to a Gamma(1, λ) prior.
As an example, the Normal-Inverse-Gamma model from Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling can be specified in PyMC with a Half-Normal prior on \(\sigma\) rather than the Inverse-Gamma:
import numpy as np
import pymc as pm
import arviz as az
# Weight measurements data from Section 5.5
rng_data = np.random.default_rng(42)
y_weights = rng_data.normal(70.5, 3.0, 15)
with pm.Model() as normal_model:
mu = pm.Normal('mu', mu=70.0, sigma=10.0)
sigma = pm.HalfNormal('sigma', sigma=5.0) # support: (0, ∞)
y = pm.Normal('y', mu=mu, sigma=sigma, observed=y_weights)
idata_norm = pm.sample(
draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False,
)
print(az.summary(idata_norm, var_names=['mu', 'sigma'], hdi_prob=0.94))
Expected output:
mean sd hdi_3% hdi_97% mcse_mean ess_bulk ess_tail r_hat
mu 70.700 0.859 69.115 72.329 0.012 5002 3216 1.00
sigma 3.208 0.625 2.141 4.335 0.008 4701 3089 1.00
The posterior mean for \(\mu\) (70.700) and \(\sigma\) (3.208) agree closely with the data mean (70.681) and sample standard deviation (3.112). The Half-Normal prior on \(\sigma\) was weakly informative enough to let the data dominate.
Common Pitfall ⚠️
Using a Normal prior on a scale parameter. A Normal(0, σ²) prior on a standard
deviation assigns probability mass to negative values, which are outside the support of σ.
PyMC will raise an error for many distributions if a parameter goes negative, or silently
return -inf log-density, stalling the chain. Always use a half-distribution or
Exponential for scale parameters. If you have an existing model from the literature that
uses a Normal prior on σ, translate it to pm.HalfNormal and note that the semantics
are slightly different (the Normal prior is effectively folded).
Derived Quantities with pm.Deterministic
Many inferential targets are not model parameters but functions of parameters. The
probability that a new patient with predictor value \(x^* = 1.5\) responds to treatment;
the ratio of two Poisson rates; the difference in group means. In PyMC, any deterministic
function of parameters can be tracked using pm.Deterministic, which stores the function’s
value at every posterior draw.
with pm.Model() as logistic_derived:
beta_0 = pm.Normal('beta_0', mu=0, sigma=2.5)
beta_1 = pm.Normal('beta_1', mu=0, sigma=2.5)
# Track derived quantities explicitly
p_at_1 = pm.Deterministic(
'P(y=1 | x=1)', pm.math.sigmoid(beta_0 + beta_1 * 1.0))
p_at_m1 = pm.Deterministic(
'P(y=1 | x=-1)', pm.math.sigmoid(beta_0 + beta_1 * (-1.0)))
odds_ratio = pm.Deterministic(
'odds_ratio', pm.math.exp(beta_1))
eta = beta_0 + beta_1 * x_raw
y = pm.Bernoulli('y', p=pm.math.sigmoid(eta), observed=y_obs)
idata_d = pm.sample(draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False)
print(az.summary(
idata_d,
var_names=['P(y=1 | x=1)', 'P(y=1 | x=-1)', 'odds_ratio'],
hdi_prob=0.94))
The pm.Deterministic values are stored in idata_d.posterior alongside the model
parameters, so az.summary and az.plot_posterior work identically for derived
quantities and parameters. This is far cleaner than post-processing the posterior array
manually and ensures that the same Monte Carlo standard errors apply.
Expected output (approximate):
mean sd hdi_3% hdi_97% r_hat
P(y=1 | x=1) 0.822 0.035 0.753 0.887 1.00
P(y=1 | x=-1) 0.161 0.031 0.104 0.220 1.00
odds_ratio 12.95 3.76 7.02 19.78 1.00
The odds ratio \(e^{\beta_1} \approx 12.95\) (94% CI: 7.02–19.78) quantifies how much the odds of a positive outcome increase per unit increase in \(x\). This is the natural inferential target for logistic regression coefficients and is obtained at zero extra computation cost once the posterior samples exist.
Practical Workflow Checklist
The following checklist consolidates the full PyMC workflow for a single-level model:
Specify the model in a
with pm.Model()block. Usepm.Datafor any predictor that may need updating for out-of-sample prediction. Wrap derived quantities inpm.Deterministic.Choose priors deliberately. Use weakly informative priors from Section 5.2 Prior Specification and Conjugate Analysis: Normal(0, 2.5²) for regression coefficients on standardised predictors; HalfNormal or HalfCauchy for scale parameters; Beta(1,1) for probability parameters.
Sample:
pm.sample(draws=2000, tune=1000, chains=4, target_accept=0.90). Increasetarget_accepttoward 0.95–0.99 if divergences appear (forces smaller step sizes).Check diagnostics — in order:
idata.sample_stats['diverging'].sum()— must be 0.az.plot_trace— visual inspection for mixing and stationarity.az.summary—r_hat ≤ 1.01, ESS ≥ 400 for all parameters.az.plot_pair— check for posterior correlations or multi-modality.
Posterior predictive check:
pm.sample_posterior_predictive, then compare simulated data distributions to observed data usingaz.plot_ppcor custom plots.Report: Posterior means, HDIs, and Monte Carlo standard errors from
az.summary. For derived quantities, usepm.Deterministicto track them automatically.
Try It Yourself 🖥️ Model Comparison Setup
The posterior samples and log-likelihoods produced in this section are the inputs to model
comparison via WAIC and LOO-CV in ch5_7-model-comparison. There are two equivalent
ways to populate idata.log_likelihood:
Option A — compute after sampling (shown throughout this section):
with logistic_model:
pm.compute_log_likelihood(idata)
with poisson_model:
pm.compute_log_likelihood(idata_pois)
# Now idata.log_likelihood['y'] is populated for both models
Option B — compute during sampling via idata_kwargs (slightly faster, avoids a
second pass through the model):
with logistic_model:
idata = pm.sample(
draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
idata_kwargs={"log_likelihood": True}, # compute during sampling
progressbar=False,
)
# idata.log_likelihood is populated immediately — no separate call needed
Both produce identical results. Option B is preferred when sampling from scratch; Option A
is convenient when you forget to set idata_kwargs or when working with an existing
idata object.
Bringing It All Together
PyMC translates the mathematical machinery of Section 5.4 Markov Chains: The Mathematical Foundation of MCMC and
Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling into a practical workflow requiring three ingredients: a model
specification written in the natural language of probability distributions, a call to
pm.sample, and an InferenceData object containing everything needed for inference,
diagnosis, and prediction. The automation is substantial — no Cholesky decompositions, no
pre-allocated innovation arrays, no acceptance rate monitoring — but the underlying machinery
is exactly what Sections 5.4 and 5.5 developed. Understanding the machinery is what makes
it possible to diagnose failures when they occur.
The two non-trivial failure modes — non-convergence flagged by r_hat > 1.01 or low ESS,
and model misspecification flagged by posterior predictive checks — each have clear remedies.
Non-convergence calls for longer chains, reparameterisation, or a better proposal (which in
PyMC means adjusting target_accept or switching to a different sampler). Model
misspecification calls for a richer likelihood, additional predictors, or a hierarchical
structure. The latter is the subject of ch5_8-hierarchical-models; model comparison
across competing specifications is the subject of ch5_7-model-comparison.
Key Takeaways 📝
Core concept: PyMC builds a symbolic computation graph from the model specification, uses Pytensor for automatic differentiation, and runs NUTS without any user-supplied gradients — the same NUTS algorithm whose theory was developed in Section 5.5 MCMC Algorithms: Metropolis-Hastings and Gibbs Sampling. (LO 4)
Computational insight: The
InferenceDataobject organises all sampling outputs — posterior samples, sample statistics, posterior predictive draws, and log-likelihoods — into a structured container; divergences inidata.sample_statsare errors, not warnings, and must be resolved before reporting results. (LO 3)Practical application: The four-step diagnostic sequence (divergences → trace plots →
az.summary→az.plot_pair) should be run before interpreting any posterior summary; posterior predictive checks assess model adequacy in a way that convergence diagnostics cannot. (LO 3, 4)Outcome alignment:
pm.Deterministicallows any function of parameters to be tracked automatically through the posterior, providing full uncertainty quantification for derived inferential targets at zero additional sampling cost. (LO 2, 4)
Exercises
Exercise 5.6.1: Probit Regression
The probit regression model replaces the logistic sigmoid with the Normal CDF:
where \(\Phi\) is the standard Normal CDF. In PyMC, this is implemented as
0.5 * (1 + pm.math.erf(eta / pm.math.sqrt(2))) using the error function from
Pytensor (which PyMC exposes via pm.math), or equivalently via
scipy.stats.norm.cdf — though the latter cannot be differentiated by PyMC’s autodiff
and would require wrapping; the pm.math.erf form is the standard approach.
(a) Fit a probit regression to the synthetic logistic dataset from Example 1. Use the same priors, number of chains, and draws.
(b) Compare az.summary output for the probit and logistic models. Are the
posterior means for \(\beta_0\) and \(\beta_1\) similar? Should they be? (Hint:
logistic and probit coefficients are on different scales — what is the approximate
conversion factor?)
(c) Run posterior predictive checks for both models. Which model produces simulated data that more closely matches the observed data?
Solution
(a)
import pymc as pm
import arviz as az
import scipy.stats as stats
with pm.Model() as probit_model:
beta_0 = pm.Normal('beta_0', mu=0, sigma=2.5)
beta_1 = pm.Normal('beta_1', mu=0, sigma=2.5)
# Probit link: Phi(eta)
eta = beta_0 + beta_1 * x_raw
p = pm.Deterministic('p', 0.5 * (1 + pm.math.erf(eta / pm.math.sqrt(2))))
y = pm.Bernoulli('y', p=p, observed=y_obs)
idata_probit = pm.sample(draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False)
pm.sample_posterior_predictive(
idata_probit, extend_inferencedata=True, random_seed=42)
(b) The probit coefficients will be approximately \(\beta_{\text{probit}} \approx \beta_{\text{logit}} / 1.7\) — the probit scale is narrower because the Normal CDF is steeper than the logistic sigmoid in the tails. For this dataset expect probit \(\beta_0 \approx -0.61\), \(\beta_1 \approx 1.51\) versus logistic \(\beta_0 \approx -1.04\), \(\beta_1 \approx 2.56\). The ratio is approximately 1.7 for both, consistent with the standard approximation.
(c) For data generated from a logistic model, the logistic posterior predictive distribution should fit marginally better, but the difference is small in practice — logistic and probit regression produce nearly identical predictions in the central probability range and differ mainly in the tails. Both models should pass the PPC with no systematic discrepancy.
Exercise 5.6.2: Negative Binomial Regression
The Poisson model assumes mean equals variance. Real count data often exhibit overdispersion — variance exceeds the mean — which the Poisson cannot capture. The Negative Binomial distribution extends Poisson with an extra dispersion parameter \(\alpha > 0\): if \(\lambda \sim \mathrm{Gamma}(\alpha, \alpha/\mu)\) and \(y \mid \lambda \sim \mathrm{Poisson}(\lambda)\), then marginally \(y \sim \mathrm{NegBin}(\mu, \alpha)\) with \(\mathrm{Var}(y) = \mu + \mu^2/\alpha\).
(a) Generate overdispersed count data:
rng = np.random.default_rng(2025)
n = 100
x = rng.normal(0, 1, n)
mu_true = np.exp(3.0 - 0.5 * x)
alpha_nb = 3.0 # dispersion; smaller = more overdispersed
y_nb = rng.negative_binomial(alpha_nb, alpha_nb / (alpha_nb + mu_true))
(b) Fit both a Poisson regression (pm.Poisson) and a Negative Binomial regression
(pm.NegativeBinomial) to this data. For the Negative Binomial use
alpha = pm.HalfNormal('alpha', sigma=5.0) as the dispersion prior.
(c) Run posterior predictive checks for both models. Which model better reproduces the variance of the observed counts? What does this tell you about the adequacy of the Poisson assumption?
Solution
(b)
# Negative Binomial model
with pm.Model() as nb_model:
a = pm.Normal('alpha_coef', mu=3.0, sigma=1.0)
b = pm.Normal('beta_coef', mu=0.0, sigma=1.0)
alpha = pm.HalfNormal('alpha', sigma=5.0)
mu_nb = pm.Deterministic('mu', pm.math.exp(a + b * x))
y_rv = pm.NegativeBinomial('y', mu=mu_nb, alpha=alpha,
observed=y_nb)
idata_nb = pm.sample(draws=2000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False)
pm.sample_posterior_predictive(
idata_nb, extend_inferencedata=True, random_seed=42)
(c) The Poisson PPC will show simulated count distributions that are too narrow — the model underestimates the variance. The Negative Binomial PPC will produce wider simulated distributions that better match the spread of the observed counts. The posterior for \(\alpha\) will be concentrated around 3 (the true value), confirming that the model correctly identifies the overdispersion level.
Exercise 5.6.3: Diagnosing a Non-Converged Model
The following model specification has a pathology that causes poor convergence:
with pm.Model() as bad_model:
mu1 = pm.Normal('mu1', mu=0, sigma=10)
mu2 = pm.Normal('mu2', mu=0, sigma=10)
# Only the sum mu1 + mu2 is identified by the data
y = pm.Normal('y', mu=mu1 + mu2, sigma=1.0,
observed=np.random.normal(5, 1, 50))
idata_bad = pm.sample(draws=2000, tune=1000, chains=4,
random_seed=42, progressbar=False)
(a) Run az.summary and az.plot_trace on idata_bad. What do r_hat and
ESS indicate?
(b) Explain in terms of the posterior geometry why this model fails to converge.
(c) Propose a reparameterisation that resolves the non-identifiability.
Solution
(a) Both mu1 and mu2 will have r_hat ≫ 1 (often 1.5–3.0) and very
low ESS (sometimes below 50). The trace plots will show the chains drifting in opposite
directions, with the sum mu1 + mu2 stable but the individual components not.
(b) The likelihood depends only on \(\mu_1 + \mu_2\), not on either individually. The posterior is a ridge — a flat direction in the \((\mu_1, \mu_2)\) space along which \(\mu_1 + \mu_2 = \text{const}\). NUTS cannot efficiently traverse this ridge because gradient information only points perpendicular to the ridge in the likelihood direction, while the prior pulls the parameters toward 0. The chain can drift along the ridge indefinitely.
(c) Reparameterise as a single sum parameter:
with pm.Model() as good_model:
mu_sum = pm.Normal('mu_sum', mu=0, sigma=14.1) # sd = sqrt(2)*10
y = pm.Normal('y', mu=mu_sum, sigma=1.0,
observed=np.random.normal(5, 1, 50))
idata_good = pm.sample(draws=2000, tune=1000, chains=4,
random_seed=42, progressbar=False)
This removes the non-identifiable decomposition entirely. In real models this arises most commonly in mixture models and hierarchical models, where the solution is to marginalise over the non-identified component or to add a hard constraint.
Exercise 5.6.4: pm.Deterministic for Risk Ratios
In a clinical trial, \(n_T = 80\) treated patients and \(n_C = 80\) control patients are enrolled. The number of responders in each group follows: \(k_T \sim \mathrm{Binomial}(n_T, \theta_T)\), \(k_C \sim \mathrm{Binomial}(n_C, \theta_C)\). Observed: \(k_T = 52\), \(k_C = 38\). Use flat Beta(1,1) priors on both rates.
(a) Fit the model in PyMC. Use pm.Deterministic to track:
The risk difference \(\Delta = \theta_T - \theta_C\)
The risk ratio \(\text{RR} = \theta_T / \theta_C\)
The odds ratio \(\text{OR} = [\theta_T/(1-\theta_T)] / [\theta_C/(1-\theta_C)]\)
(b) Report the posterior means and 94% HDIs for all three effect measures.
(c) What is the posterior probability that the treatment is beneficial (\(P(\theta_T > \theta_C \mid y)\))?
Solution
(a) and (b):
import pymc as pm
import arviz as az
n_T, k_T = 80, 52
n_C, k_C = 80, 38
with pm.Model() as rct_model:
theta_T = pm.Beta('theta_T', alpha=1, beta=1)
theta_C = pm.Beta('theta_C', alpha=1, beta=1)
delta = pm.Deterministic('risk_difference', theta_T - theta_C)
rr = pm.Deterministic('risk_ratio', theta_T / theta_C)
or_ = pm.Deterministic(
'odds_ratio',
(theta_T / (1 - theta_T)) / (theta_C / (1 - theta_C)))
y_T = pm.Binomial('y_T', n=n_T, p=theta_T, observed=k_T)
y_C = pm.Binomial('y_C', n=n_C, p=theta_C, observed=k_C)
idata_rct = pm.sample(draws=4000, tune=1000, chains=4,
target_accept=0.90, random_seed=42,
progressbar=False)
print(az.summary(
idata_rct,
var_names=['theta_T', 'theta_C',
'risk_difference', 'risk_ratio', 'odds_ratio'],
hdi_prob=0.94))
Expected output (approximate):
mean sd hdi_3% hdi_97%
theta_T 0.645 0.052 0.545 0.742
theta_C 0.481 0.054 0.374 0.579
risk_difference 0.164 0.074 0.022 0.306
risk_ratio 1.350 0.171 1.036 1.674
odds_ratio 2.042 0.569 1.118 3.100
(c)
delta_samples = idata_rct.posterior['risk_difference'].values.flatten()
p_beneficial = (delta_samples > 0).mean()
print(f"P(θ_T > θ_C | y) = {p_beneficial:.4f}")
# Expected: approximately 0.986
References
PyMC and ArviZ
Salvatier, J., Wiecki, T. V., and Fonnesbeck, C. (2016). Probabilistic programming in Python using PyMC3. PeerJ Computer Science, 2, e55. The original PyMC3 paper; the API has evolved through PyMC4 and PyMC 5 but the probabilistic programming philosophy described here remains the same.
PyMC Development Team (2023). PyMC: A modern and comprehensive probabilistic programming framework in Python. Version 5. Available at: https://www.pymc.io. The software library providing all PyMC functionality used in this section.
Kumar, R., Carroll, C., Hartikainen, A., and Martin, O. (2019). ArviZ a unified library for exploratory analysis of Bayesian models in Python. Journal of Open Source Software, 4(33), 1143. The ArviZ library providing az.summary, az.plot_trace, az.plot_pair, az.plot_ppc, and all other diagnostic visualisations used in this section.
Model Specification and Priors
Gelman, A. (2006). Prior distributions for variance parameters in hierarchical models. Bayesian Analysis, 1(3), 515–534. The source of the Half-Cauchy(0, 2.5) recommendation for hierarchical variance components; widely adopted in PyMC and Stan communities.
Gelman, A., and Hill, J. (2007). Data Analysis Using Regression and Multilevel/Hierarchical Models, Ch. 16. Cambridge University Press. The Gelman recommendation for Normal(0, 2.5²) weakly informative priors for logistic regression coefficients on standardised predictors.
Diagnostics
Vehtari, A., Gelman, A., Simpson, D., Carpenter, B., and Bürkner, P.-C. (2021). Rank-normalization, folding, and localization: An improved \(\hat{R}\) for assessing convergence of MCMC. Bayesian Analysis, 16(2), 667–718. The rank-normalised \(\hat{R}\) and ESS diagnostics implemented in ArviZ and explained from first principles in Section 5.4 Markov Chains: The Mathematical Foundation of MCMC.
Betancourt, M. (2017). A conceptual introduction to Hamiltonian Monte Carlo. arXiv:1701.02434. An accessible but rigorous treatment of HMC, NUTS, and the diagnostic significance of divergences; the basis for understanding why divergences cannot be ignored.
Posterior Predictive Checks
Gelman, A., Meng, X.-L., and Stern, H. (1996). Posterior predictive assessment of model fitness via realized discrepancies. Statistica Sinica, 6(4), 733–807. The foundational paper on posterior predictive checks; defines the posterior predictive p-value as a measure of model fit.
Gelman, A., Carlin, J. B., Stern, H. S., Dunson, D. B., Vehtari, A., and Rubin, D. B. (2013). Bayesian Data Analysis (3rd ed.), Ch. 6. Chapman and Hall/CRC. Standard treatment of posterior predictive checks and calibration assessment; the az.plot_ppc function implements the visualisations from this chapter.